lattpy.spatial
Spatial algorithms and data structures.
- lattpy.spatial.distance(r1, r2, decimals=None)[source]
Calculates the euclidian distance bewteen two points.
- Parameters
- r1(D, ) np.ndarray
First input point.
- r2(D, ) np.ndarray
Second input point of matching size.
- decimals: int, optional
Decimals for rounding the output.
- Returns
- distance: float
The distance between the input points.
- lattpy.spatial.distances(r1, r2, decimals=None)[source]
Calculates the euclidian distance between multiple points.
- Parameters
- r1(N, D) array_like
First input point.
- r2(N, D) array_like
Second input point of matching size.
- decimals: int, optional
Optional decimals to round distance to.
- Returns
- distances(N, ) np.ndarray
The distances for each pair of input points.
- lattpy.spatial.interweave(arrays)[source]
Interweaves multiple arrays along the first axis
- Parameters
- arrays: (M) Sequence of (N, …) array_like
The input arrays to interwave. The shape of all arrays must match.
- Returns
- interweaved: (M*N, ….) np.ndarray
- lattpy.spatial.vindices(limits, sort_axis=0, dtype=None)[source]
Return an array representing the indices of a d-dimensional grid.
- Parameters
- limits: (D, 2) array_like
The limits of the indices for each axis.
- sort_axis: int, optional
Optional axis that is used to sort indices.
- dtype: int or str or np.dtype, optional
Optional data-type for storing the lattice indices. By default the given limits are checked to determine the smallest possible data-type.
- Returns
- vectors: (N, D) np.ndarray
- lattpy.spatial.vrange(start=None, *args, dtype=None, sort_axis=0, **kwargs)[source]
Return evenly spaced vectors within a given interval.
- Parameters
- start: array_like, optional
The starting value of the interval. The interval includes this value. The default start value is 0.
- stop: array_like
The end value of the interval.
- step: array_like, optional
Spacing between values. If start and stop are sequences and the step is a scalar the given step size is used for all dimensions of the vectors. The default step size is 1.
- sort_axis: int, optional
Optional axis that is used to sort indices.
- dtype: dtype, optional
The type of the output array. If dtype is not given, infer the data type from the other input arguments.
- Returns
- vectors: (N, D) np.ndarray
- lattpy.spatial.cell_size(vectors)[source]
Computes the shape of the box spawned by the given vectors.
- Parameters
- vectors: array_like
The basis vectors defining the cell.
- Returns
- size: np.ndarray
- lattpy.spatial.cell_volume(vectors)[source]
Computes the volume of the unit cell defined by the primitive vectors.
The volume of the unit-cell in two and three dimensions is defined by
\[V_{2d} = \abs{a_1 \cross a_2}, \quad V_{3d} = a_1 \cdot \abs{a_2 \cross a_3}\]For higher dimensions the volume is computed using the determinant:
\[V_{d} = \sqrt{\det{A A^T}}\]where \(A\) is the array of vectors.
- Parameters
- vectors: array_like
The basis vectors defining the cell.
- Returns
- vol: float
The volume of the unit cell.
- lattpy.spatial.compute_vectors(a, b=None, c=None, alpha=None, beta=None, gamma=None, decimals=0)[source]
Computes lattice vectors by the lengths and angles.
- class lattpy.spatial.KDTree(points, k=1, max_dist=inf, eps=0.0, p=2, boxsize=None)[source]
Bases:
cKDTreeSimple wrapper of scipy’s cKTree with global query settings.
- query_ball_point(self, x, r, p=2.0, eps=0, workers=1, return_sorted=None, return_length=False)[source]
Find all points within distance r of point(s) x.
- Parameters
- xarray_like, shape tuple + (self.m,)
The point or points to search for neighbors of.
- rarray_like, float
The radius of points to return, shall broadcast to the length of x.
- pfloat, optional
Which Minkowski p-norm to use. Should be in the range [1, inf]. A finite large p may cause a ValueError if overflow can occur.
- epsnonnegative float, optional
Approximate search. Branches of the tree are not explored if their nearest points are further than
r / (1 + eps), and branches are added in bulk if their furthest points are nearer thanr * (1 + eps).- workersint, optional
Number of jobs to schedule for parallel processing. If -1 is given all processors are used. Default: 1.
Changed in version 1.6.0: The “n_jobs” argument was renamed “workers”. The old name “n_jobs” is deprecated and will stop working in SciPy 1.8.0.
- return_sortedbool, optional
Sorts returned indicies if True and does not sort them if False. If None, does not sort single point queries, but does sort multi-point queries which was the behavior before this option was added.
New in version 1.2.0.
- return_length: bool, optional
Return the number of points inside the radius instead of a list of the indices. .. versionadded:: 1.3.0
- Returns
- resultslist or array of lists
If x is a single point, returns a list of the indices of the neighbors of x. If x is an array of points, returns an object array of shape tuple containing lists of neighbors.
Notes
If you have many points whose neighbors you want to find, you may save substantial amounts of time by putting them in a cKDTree and using query_ball_tree.
Examples
>>> from scipy import spatial >>> x, y = np.mgrid[0:4, 0:4] >>> points = np.c_[x.ravel(), y.ravel()] >>> tree = spatial.cKDTree(points) >>> tree.query_ball_point([2, 0], 1) [4, 8, 9, 12]
Query multiple points and plot the results:
>>> import matplotlib.pyplot as plt >>> points = np.asarray(points) >>> plt.plot(points[:,0], points[:,1], '.') >>> for results in tree.query_ball_point(([2, 0], [3, 3]), 1): ... nearby_points = points[results] ... plt.plot(nearby_points[:,0], nearby_points[:,1], 'o') >>> plt.margins(0.1, 0.1) >>> plt.show()

- query_ball_tree(self, other, r, p=2.0, eps=0)[source]
Find all pairs of points between self and other whose distance is at most r
- Parameters
- othercKDTree instance
The tree containing points to search against.
- rfloat
The maximum distance, has to be positive.
- pfloat, optional
Which Minkowski norm to use. p has to meet the condition
1 <= p <= infinity. A finite large p may cause a ValueError if overflow can occur.- epsfloat, optional
Approximate search. Branches of the tree are not explored if their nearest points are further than
r/(1+eps), and branches are added in bulk if their furthest points are nearer thanr * (1+eps). eps has to be non-negative.
- Returns
- resultslist of lists
For each element
self.data[i]of this tree,results[i]is a list of the indices of its neighbors inother.data.
Examples
You can search all pairs of points between two kd-trees within a distance:
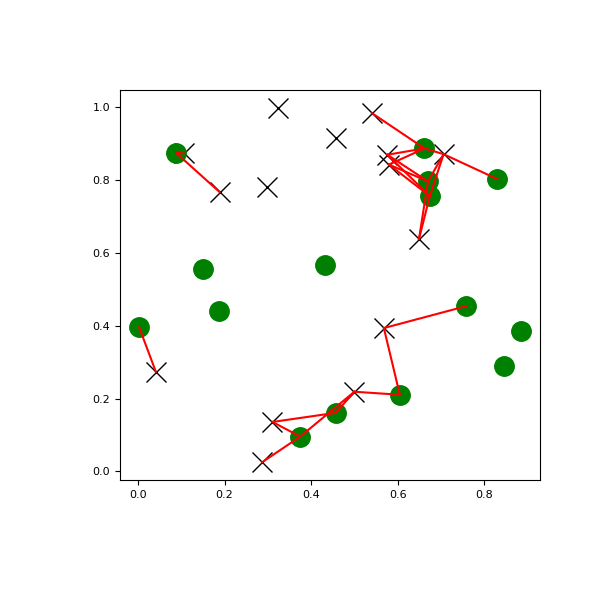
>>> import matplotlib.pyplot as plt >>> import numpy as np >>> from scipy.spatial import cKDTree >>> rng = np.random.default_rng() >>> points1 = rng.random((15, 2)) >>> points2 = rng.random((15, 2)) >>> plt.figure(figsize=(6, 6)) >>> plt.plot(points1[:, 0], points1[:, 1], "xk", markersize=14) >>> plt.plot(points2[:, 0], points2[:, 1], "og", markersize=14) >>> kd_tree1 = cKDTree(points1) >>> kd_tree2 = cKDTree(points2) >>> indexes = kd_tree1.query_ball_tree(kd_tree2, r=0.2) >>> for i in range(len(indexes)): ... for j in indexes[i]: ... plt.plot([points1[i, 0], points2[j, 0]], ... [points1[i, 1], points2[j, 1]], "-r") >>> plt.show()
- query_pairs(self, r, p=2.0, eps=0)[source]
Find all pairs of points in self whose distance is at most r.
- Parameters
- rpositive float
The maximum distance.
- pfloat, optional
Which Minkowski norm to use.
phas to meet the condition1 <= p <= infinity. A finite large p may cause a ValueError if overflow can occur.- epsfloat, optional
Approximate search. Branches of the tree are not explored if their nearest points are further than
r/(1+eps), and branches are added in bulk if their furthest points are nearer thanr * (1+eps). eps has to be non-negative.- output_typestring, optional
Choose the output container, ‘set’ or ‘ndarray’. Default: ‘set’
- Returns
- resultsset or ndarray
Set of pairs
(i,j), withi < j, for which the corresponding positions are close. If output_type is ‘ndarray’, an ndarry is returned instead of a set.
Examples
You can search all pairs of points in a kd-tree within a distance:
>>> import matplotlib.pyplot as plt >>> import numpy as np >>> from scipy.spatial import cKDTree >>> rng = np.random.default_rng() >>> points = rng.random((20, 2)) >>> plt.figure(figsize=(6, 6)) >>> plt.plot(points[:, 0], points[:, 1], "xk", markersize=14) >>> kd_tree = cKDTree(points) >>> pairs = kd_tree.query_pairs(r=0.2) >>> for (i, j) in pairs: ... plt.plot([points[i, 0], points[j, 0]], ... [points[i, 1], points[j, 1]], "-r") >>> plt.show()

- query(self, x, k=1, eps=0, p=2, distance_upper_bound=np.inf, workers=1)[source]
Query the kd-tree for nearest neighbors
- Parameters
- xarray_like, last dimension self.m
An array of points to query.
- klist of integer or integer
The list of k-th nearest neighbors to return. If k is an integer it is treated as a list of [1, … k] (range(1, k+1)). Note that the counting starts from 1.
- epsnon-negative float
Return approximate nearest neighbors; the k-th returned value is guaranteed to be no further than (1+eps) times the distance to the real k-th nearest neighbor.
- pfloat, 1<=p<=infinity
Which Minkowski p-norm to use. 1 is the sum-of-absolute-values “Manhattan” distance 2 is the usual Euclidean distance infinity is the maximum-coordinate-difference distance A finite large p may cause a ValueError if overflow can occur.
- distance_upper_boundnonnegative float
Return only neighbors within this distance. This is used to prune tree searches, so if you are doing a series of nearest-neighbor queries, it may help to supply the distance to the nearest neighbor of the most recent point.
- workersint, optional
Number of workers to use for parallel processing. If -1 is given all CPU threads are used. Default: 1.
Changed in version 1.6.0: The “n_jobs” argument was renamed “workers”. The old name “n_jobs” is deprecated and will stop working in SciPy 1.8.0.
- Returns
- darray of floats
The distances to the nearest neighbors. If
xhas shapetuple+(self.m,), thendhas shapetuple+(k,). When k == 1, the last dimension of the output is squeezed. Missing neighbors are indicated with infinite distances.- indarray of ints
The index of each neighbor in
self.data. Ifxhas shapetuple+(self.m,), thenihas shapetuple+(k,). When k == 1, the last dimension of the output is squeezed. Missing neighbors are indicated withself.n.
Notes
If the KD-Tree is periodic, the position
xis wrapped into the box.When the input k is a list, a query for arange(max(k)) is performed, but only columns that store the requested values of k are preserved. This is implemented in a manner that reduces memory usage.
Examples
>>> import numpy as np >>> from scipy.spatial import cKDTree >>> x, y = np.mgrid[0:5, 2:8] >>> tree = cKDTree(np.c_[x.ravel(), y.ravel()])
To query the nearest neighbours and return squeezed result, use
>>> dd, ii = tree.query([[0, 0], [2.2, 2.9]], k=1) >>> print(dd, ii, sep='\n') [2. 0.2236068] [ 0 13]
To query the nearest neighbours and return unsqueezed result, use
>>> dd, ii = tree.query([[0, 0], [2.2, 2.9]], k=[1]) >>> print(dd, ii, sep='\n') [[2. ] [0.2236068]] [[ 0] [13]]
To query the second nearest neighbours and return unsqueezed result, use
>>> dd, ii = tree.query([[0, 0], [2.2, 2.9]], k=[2]) >>> print(dd, ii, sep='\n') [[2.23606798] [0.80622577]] [[ 6] [19]]
To query the first and second nearest neighbours, use
>>> dd, ii = tree.query([[0, 0], [2.2, 2.9]], k=2) >>> print(dd, ii, sep='\n') [[2. 2.23606798] [0.2236068 0.80622577]] [[ 0 6] [13 19]]
or, be more specific
>>> dd, ii = tree.query([[0, 0], [2.2, 2.9]], k=[1, 2]) >>> print(dd, ii, sep='\n') [[2. 2.23606798] [0.2236068 0.80622577]] [[ 0 6] [13 19]]
- class lattpy.spatial.WignerSeitzCell(points)[source]
Bases:
VoronoiTree- property limits
- property size